Folding Farm
For a while I got quite interested in the Folding@Home project, where you use home computers to figure out the way certain proteins are folded in order to aid medical research. It’s a way for medical researchers to utilise all the untapped computing power in people’s desktop computers all over the world.
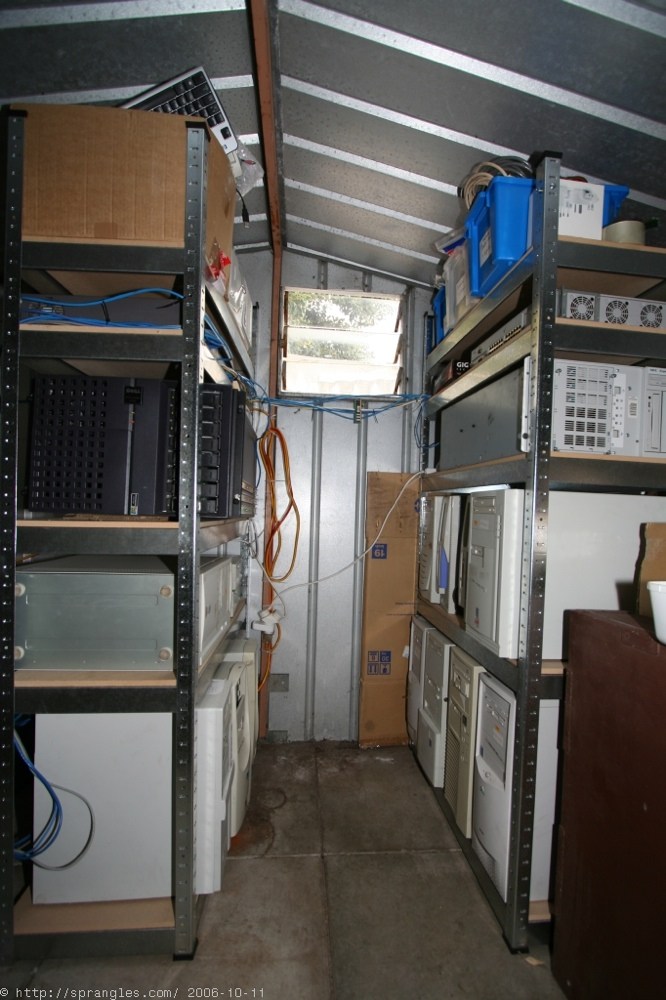
I took it a step further by collecting old computers and networking them in to a mini-datacentre in my shed (though “datacentre” is overselling it a bit). I tried to install a different operating system on each one (different flavours of Linux, FreeBSD and Windows) so that I would have a bit of experience with each one and better understand the different environments. I ended up with 22 hosts in total, busily folding away at proteins. It became a bit of a challenge to see if I could pick up some decent hardware for less than $50 through the used-PC classifieds.
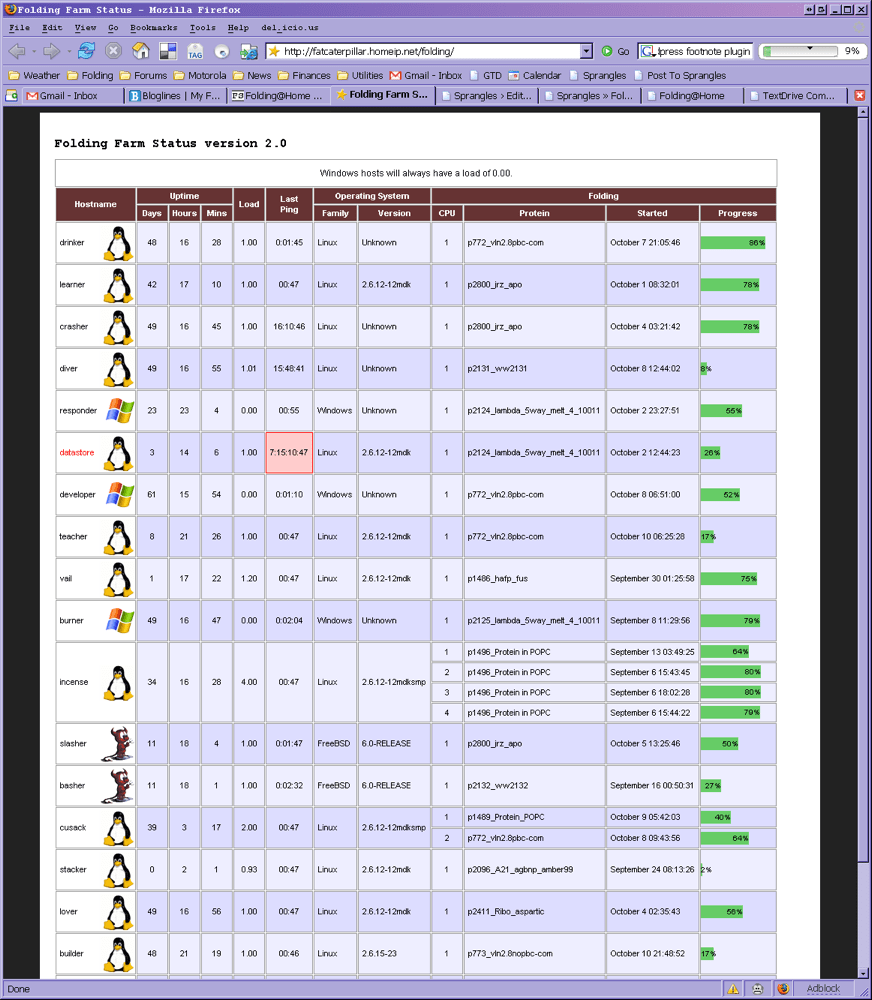
I wrote some monitoring software to keep track of all the machines: which protein they were working on, their current status and how long they’d been running. It was quite satisfying to think of all that work being done 24/7 in my shed.
In the end I decommissioned the farm and chucked out all the machines for a few reasons:
- standard computing power crept forward so that my farm was really behind the curve – I was no longer getting enough computing power for the running costs relative to what everyone was able to achieve (my contribution was getting relatively smaller and smaller each day)
- I wanted to use my shed for other things!
I encourage you to use your computers to do some folding though, you don’t need a big cluster to be useful. You can use your computer’s spare cycles tohelp find cures for diseases like Parkinson’s and Alzheimer’s.